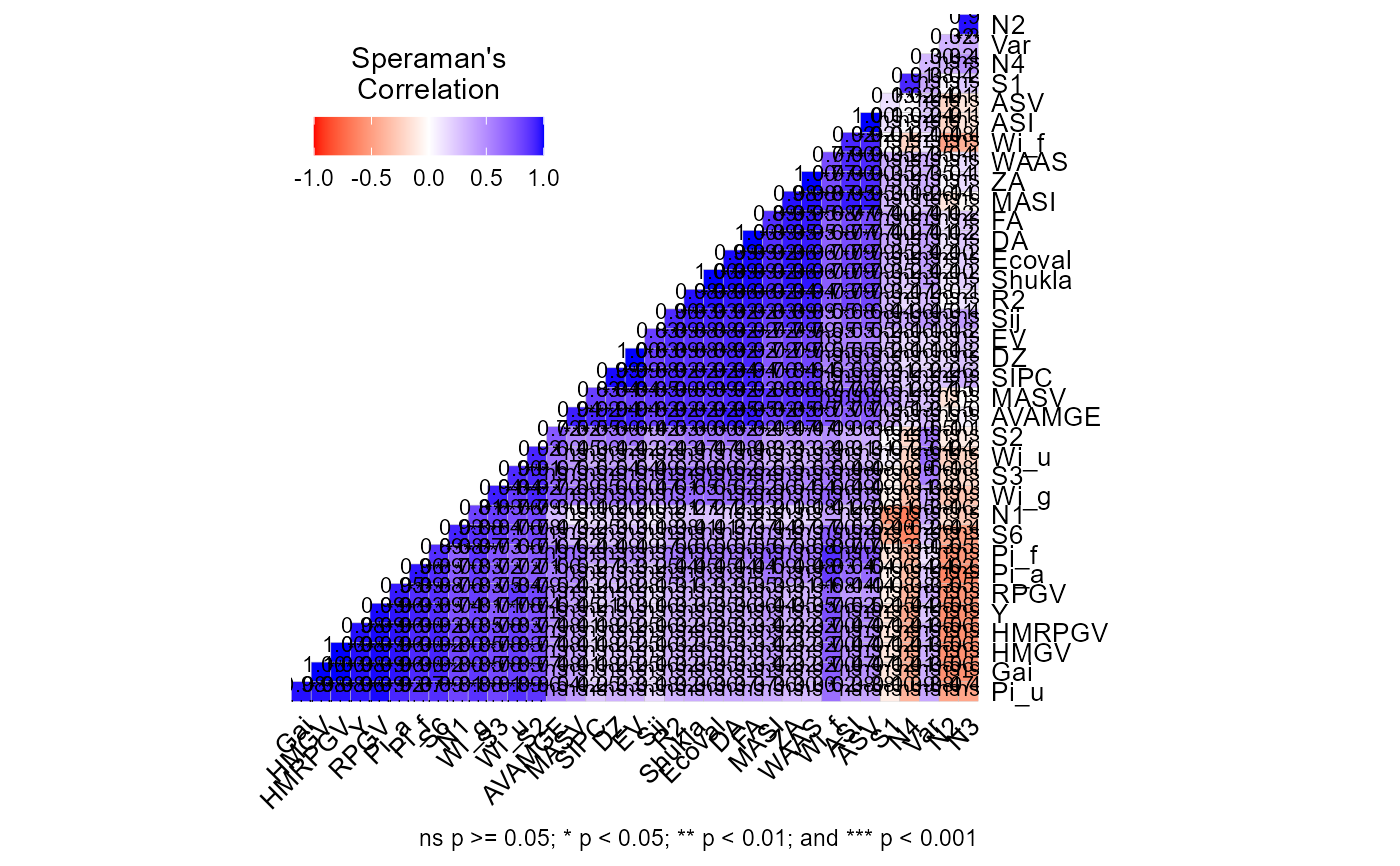
Computes the Spearman's rank correlation between the parametric and
nonparametric stability indexes computed with the function
ge_stats().
Arguments
- x
An object of class
ge_stats.- stats
The statistics to compute the correlation. See the section Details for more information.
- plot
Plot the heat map with the correlations? Defaults to
TRUE.- ...
Other arguments to be passed to the function
plot.corr_coef().
Value
A list with the data (ranks) correlation, p-values and a heat map showing the correlation coefficients.
Details
The argument stats is used to chose the statistics to show the
ranks. Allowed values are "all" (All statistics, default), "par"
(Parametric statistics), "nonpar" (Non-parametric statistics), "ammi"
(AMMI-based stability statistics), or the following values that can be
combined into comma-separated character vector. "Y" (Response variable),
"Var" (Genotype's variance), "Shukla" (Shukla's variance), "Wi_g", "Wi_f", "Wi_u" (Annichiarrico's genotypic confidence index for all,
favorable and unfavorable environments, respectively), "Ecoval" (Wricke's
ecovalence), "Sij" (Deviations from the joint-regression analysis),
"R2" (R-squared from the joint-regression analysis), "ASTAB" (AMMI
Based Stability Parameter), "ASI" (AMMI Stability Index), "ASV"
(AMMI-stability value), "AVAMGE" (Sum Across Environments of Absolute
Value of GEI Modelled by AMMI ), "Da" (Annicchiarico's D Parameter
values), "Dz" (Zhang's D Parameter), "EV" (Sums of the Averages of the
Squared Eigenvector Values), "FA" (Stability Measure Based on Fitted AMMI
Model), "MASV" (Modified AMMI Stability Value), "SIPC" (Sums of the
Absolute Value of the IPC Scores), "Za" (Absolute Value of the Relative
Contribution of IPCs to the Interaction), "WAAS" (Weighted average of
absolute scores), "HMGV" (Harmonic mean of the genotypic value), "RPGV"
(Relative performance of the genotypic values), "HMRPGV" (Harmonic mean
of the relative performance of the genotypic values), "Pi_a", "Pi_f", "Pi_u" (Superiority indexes for all, favorable and unfavorable
environments, respectively), "Gai" (Geometric adaptability index), "S1"
(mean of the absolute rank differences of a genotype over the n
environments), "S2" (variance among the ranks over the k environments),
"S3" (sum of the absolute deviations), "S6" (relative sum of squares of
rank for each genotype), "N1", "N2", "N3", "N4" (Thennarasu"s
statistics)).
Author
Tiago Olivoto tiagoolivoto@gmail.com