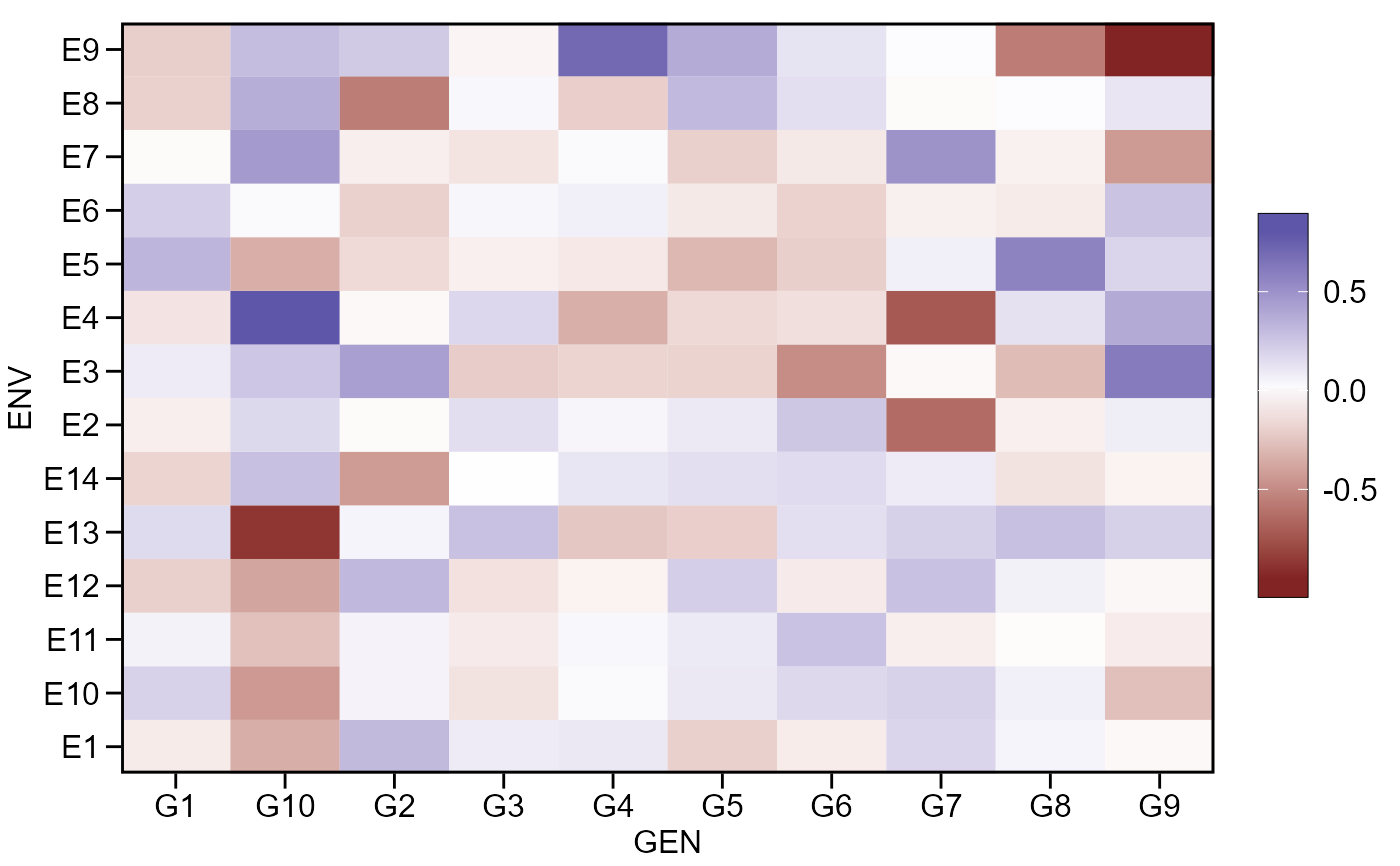
This is a helper function that computes the genotype-environment effects, i.e., the residual effect of the additive model
Arguments
- .data
The dataset containing the columns related to Environments, Genotypes, replication/block and response variable(s).
- env
The name of the column that contains the levels of the environments. The analysis of variance is computed for each level of this factor.
- gen
The name of the column that contains the levels of the genotypes.
- resp
The response variable(s). To analyze multiple variables in a single procedure a vector of variables may be used. For example
resp = c(var1, var2, var3).- type
The type of effect to compute. Defaults to
"ge", i.e., genotype-environment. To compute genotype plus genotype-environment effects usetype = "gge".- verbose
Logical argument. If
verbose = FALSEthe code will run silently.
Value
A list where each element is the result for one variable that contains a two-way table with genotypes in rows and environments in columns.
Author
Tiago Olivoto tiagoolivoto@gmail.com