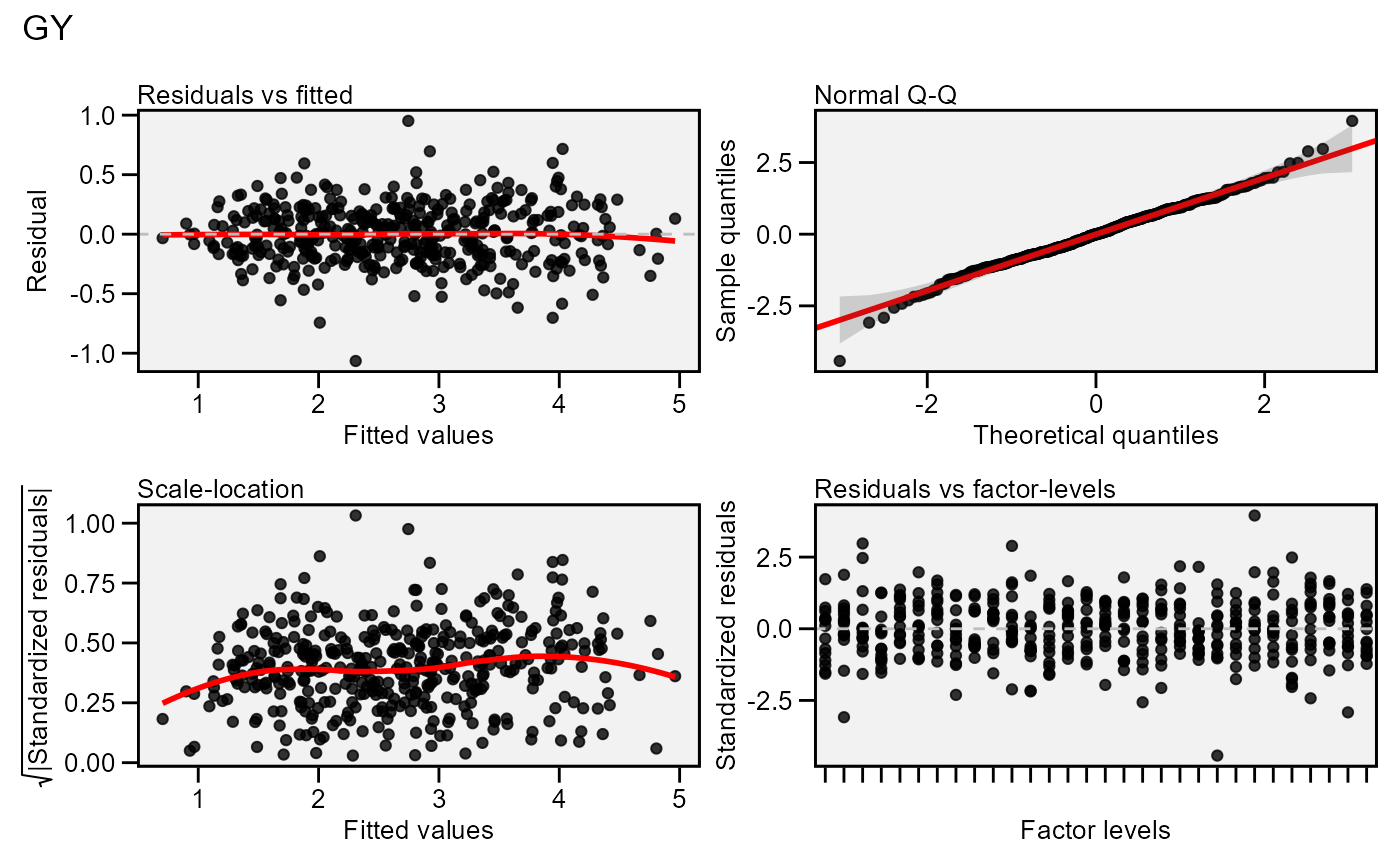
Residual plots for a output model of class performs_ammi. Seven types
of plots are produced: (1) Residuals vs fitted, (2) normal Q-Q plot for the
residuals, (3) scale-location plot (standardized residuals vs Fitted Values),
(4) standardized residuals vs Factor-levels, (5) Histogram of raw residuals
and (6) standardized residuals vs observation order, and (7) 1:1 line plot.
Usage
# S3 method for performs_ammi
plot(x, ...)Arguments
- x
An object of class
performs_ammi.- ...
Additional arguments passed on to the function
residual_plots()
Author
Tiago Olivoto tiagoolivoto@gmail.com
Examples
# \donttest{
library(metan)
model <- performs_ammi(data_ge, ENV, GEN, REP, GY)
#> variable GY
#> ---------------------------------------------------------------------------
#> AMMI analysis table
#> ---------------------------------------------------------------------------
#> Source Df Sum Sq Mean Sq F value Pr(>F) Proportion Accumulated
#> ENV 13 279.574 21.5057 62.33 0.00e+00 NA NA
#> REP(ENV) 28 9.662 0.3451 3.57 3.59e-08 NA NA
#> GEN 9 12.995 1.4439 14.93 2.19e-19 NA NA
#> GEN:ENV 117 31.220 0.2668 2.76 1.01e-11 NA NA
#> PC1 21 10.749 0.5119 5.29 0.00e+00 34.4 34.4
#> PC2 19 9.924 0.5223 5.40 0.00e+00 31.8 66.2
#> PC3 17 4.039 0.2376 2.46 1.40e-03 12.9 79.2
#> PC4 15 3.074 0.2049 2.12 9.60e-03 9.8 89.0
#> PC5 13 1.446 0.1113 1.15 3.18e-01 4.6 93.6
#> PC6 11 0.932 0.0848 0.88 5.61e-01 3.0 96.6
#> PC7 9 0.567 0.0630 0.65 7.53e-01 1.8 98.4
#> PC8 7 0.362 0.0518 0.54 8.04e-01 1.2 99.6
#> PC9 5 0.126 0.0252 0.26 9.34e-01 0.4 100.0
#> Residuals 252 24.367 0.0967 NA NA NA NA
#> Total 536 389.036 0.7258 NA NA NA NA
#> ---------------------------------------------------------------------------
#>
#> All variables with significant (p < 0.05) genotype-vs-environment interaction
#> Done!
plot(model)
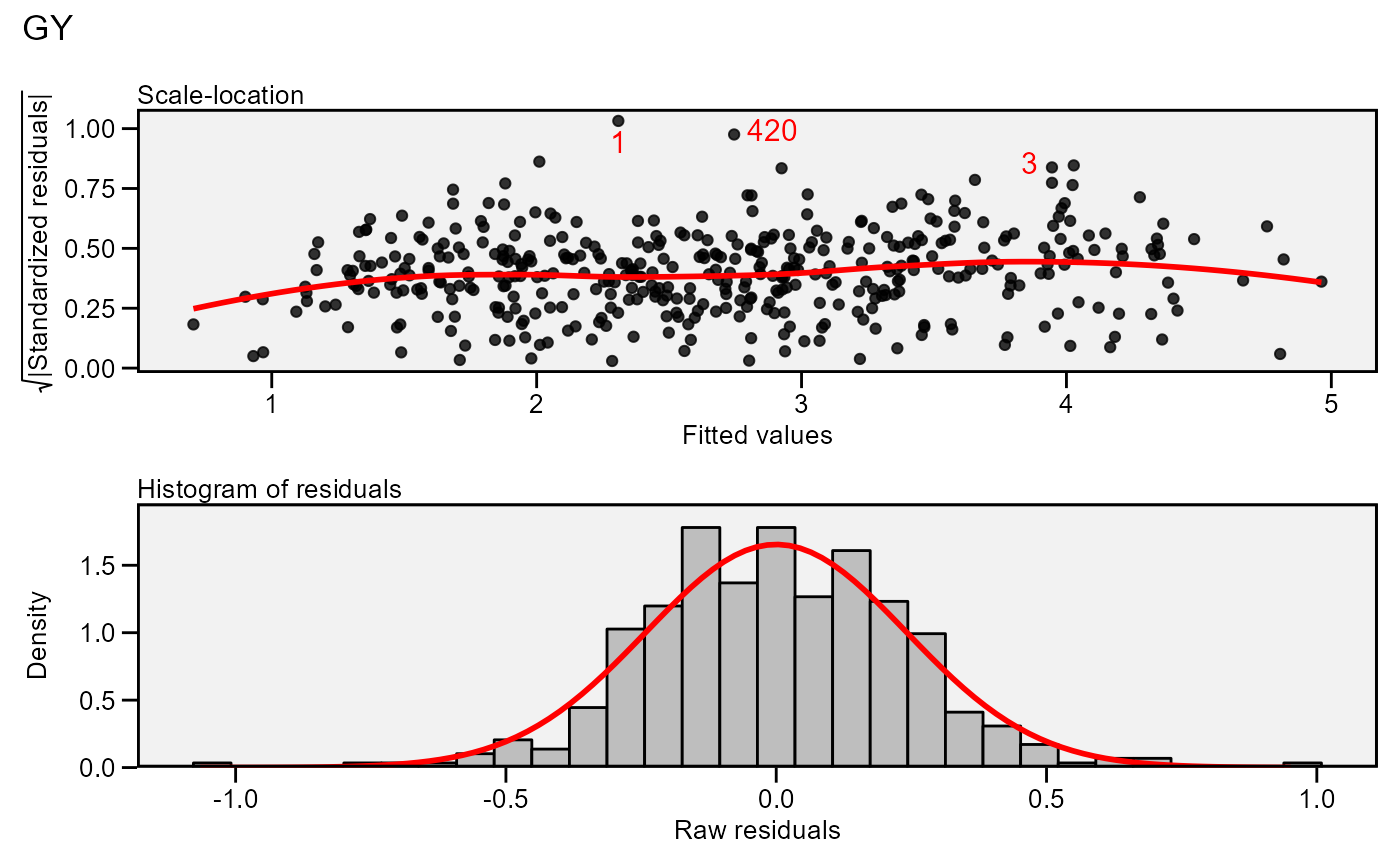
 plot(model,
which = c(3, 5),
nrow = 2,
labels = TRUE,
size.lab.out = 4)
plot(model,
which = c(3, 5),
nrow = 2,
labels = TRUE,
size.lab.out = 4)
 # }
# }
